Publications
Shoemaker S, Luo M, Dam KM, Pak J, Hoffmann M, Marqusee S. (2025) Conformational dynamics of SARS-CoV-2 spike on a membrane reveals the allosteric effects of furin cleavage and the D614G mutation. Science Advances. DOI: doi.org/10.1126/sciadv.adx51786 [link] [bioRxiv]
Williams C, Nocka L, Hedger G, Parashara P, Pardon E, Latorraca N, Pusapati G, Lartey D, Gao L, Milenkovic L, Chalk R, Steyaert J, Marqqusee S, Carrique L, Bazan JF, Rouse S, Kong J, Siebold C, Rohatgi R. (2025) Design principles of a membrane-spanning ubiquitin ligase. BioRxiv. DOI: doi.org/10.1101/2025.09.11.675358 [link]
Hood M, Marqusee S. (2025) Exploring the sequence and structural determinants of the energy landscape from thermodynamically stable and kinetically trapped subtilisins: ISP1 and SbtE. Protein Sci. DOI: doi.org/10.1002/pro.70264 [link] [bioRxiv]
De Cae S, Van Molle I, van Schie L, Shoemaker S, Deckers J, Debeuf N, Lameire S, Nerinckx W, Roose K, Fijalkowska D, Devos S, De Smet AS, Marchan JCZ, Venneman T, Sedeyn K, Mujanovic L, Ballegeer M, Vanheerswynghels M, De Wolf C, Demol H, Zuallaert J, Vanhaverbeke P, Ghassabeh GH, Lonigro C, Bockstal V, Rinaldi M, Abdelnabi R, Neyts J, Marqusee S, Lambrecht BN, Callewaert N, Remaut H, Saelens X, Schepens B. (2025) Ultrapotent SARS coronavirus-neutralizing single-domain antibodies that clamp the spike at its base. Nat Commun. DOI: doi.org/10.1038/s41467-025-60250-1 [link]
Latorraca N, Sabaat S, Habrian C, Bleier J, Stanley C, Kinz-Thompson C, Marqusee S, Isacoff E. (2025) Domain coupling in activation of a family C GPCR. Nat Chem Biol. DOI: doi.org/10.1038/s41589-025-01895-3 [link] [bioRxiv]
Arkinson C, Dong KC, Gee CL, Costello SM, Soe AC, Hura GL, Marqusee S, Martin A. (2025) Nub1 traps unfolded FAT10 for ubiquitin-independent degradation by the 26S proteasome. Nat Struct Mol Biol. DOI: doi.org/10.1038/s41594-025-01527-3 [link] [bioRxiv]
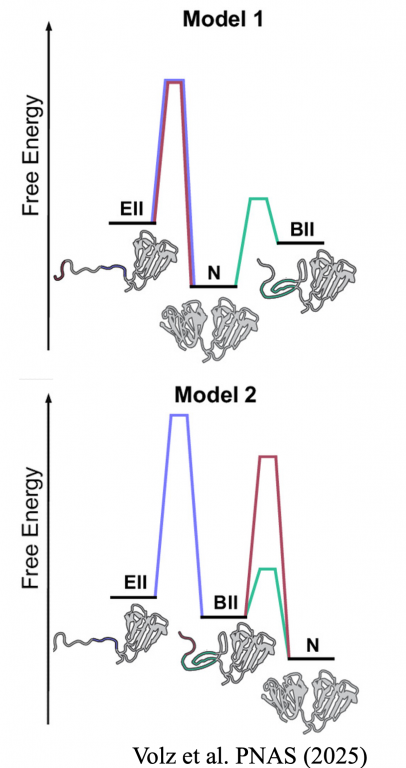
Volz S, Malone JR, Guseman AJ, Gronenborn AM, Marqusee S. (2025) Cateract-prone variants of γD-crystallin populate a conformation with a partially unfolded N-terminal domain under native conditions. Proc Natl Acad Sci USA. DOI: doi.org/10.1073/pnas.2410860122 [link]
Gupta S, Russell B, Kristensen LG, Tyler J, Costello SM, Marqusee S, Rad B, Ralston C. (2025) Enabling simultaneous photoluminescence spectroscopy and X-ray footprinting mass spectrometry to study protein conformation and interactions. Anal Methods. DOI: doi.org/10.1039/d4ay01670j [link]
Dall NR, Mendonça CATF, Torres Vera HL, Marqusee S. (2024) The importance of the location of the N-terminus in successful protein folding in vivo and in vitro. Proc Natl Acad Sci USA. DOI: doi.org/10.1073/pnas.2321999121 [link]
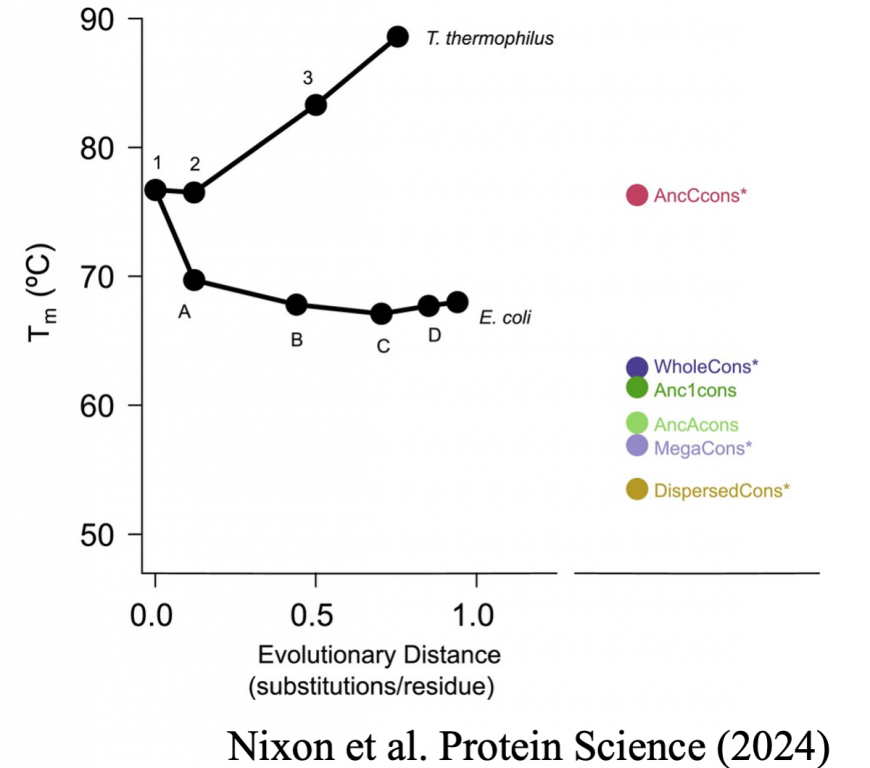
Nixon C, Lim S, Sternke M, Barrick D, Harms M, Marqusee S. (2024) The importance of input sequence set to consensus-derived proteins and their relationship to reconstructed ancestral proteins. Protein Science. DOI: doi.org/10.1002/pro.5011 [link] [bioRxiv]
Kumar KK, Wang H, Habrian C, Latorraca N, Xu J, O'Brien E, Zhang C, Montabana E, Koehl A, Marqusee S, Isacoff E, Kobilka B. (2024) Stepwise activation of a metabotropic glutamate receptor. Nature. DOI: doi.org/10.1038/s41586-024-07327-x [link]
Hayes RL, Nixon CF, Marqusee S, Brooks CL 3rd. (2024) Selection pressures on evolution of ribonuclease H explored with rigorous free-energy-based design. Proc Natl Acad Sci USA. DOI: doi.org/10.1073/pnas.2312029121 [link]
Gupta S, Inman JL, Chant J, Obst-Huebl L, Nakamura K, Costello SM, Marqusee S, Mao JH, Kunz L, Paisley R, Vozenin MC, Snijders AM, Ralston CY. (2023) A novel platform for evaluating dose rate effects on oxidative damage to peptides: toward a high-throughput method to characterize the mechanisms underlying the FLASH effect. Radiat Res. DOI: doi.org/10.1667/RADE-23-00131.1 [link]
Lawrence R, Shoemaker S, Deal A, Sangwan S, Anand A, Wang L, Marqusee S, Walter P. (2023) A helical fulcrum in eIF2B coordinates allosteric regulation of stress signaling. Nat Chem Biol. DOI: doi.org/10.1038/s41589-023-01453-9 [link] [bioRxiv]
De Cae S, Van Molle I, van Schie L, Shoemaker SR, Deckers J, Debeuf N, Lameire S, Nerinckx W, Roose K, Fijalkowska D, Devos S, Desmet AS, Marchan JCZ, Venneman T, Sedeyn K, Ballegeer M, Vanheerswynghels M, De Wolf C, Demol H, Vanhaverbeke P, Ghassabeh GH, Lonigro C, Bockstal V, Rinaldi M, Abdelnabi R, Neyts J, Marqusee S, Lambrecht BN, Callewaert N, Remaut H, Saelens X, Schepens B. (2023) Ultrapotent SARS coronavirus-neutralizing single-domain antibodies that bind a conserved membrane proximal epitope of the spike. bioRxiv. DOI: doi.org/10.1101/2023.03.10.531533 [bioRxiv]
Shaw A, Craig JM, Amiri H, Kim J, Upton HE, Pimentel SC, Huang JR, Marqusee S, Gundlach JH, Collins K, Bustamante CJ. (2023) Secondary structure detection through direct nanopore RNA sequencing. bioRxiv. DOI: doi.org/10.1101/2023.04.05.535757 [bioRxiv]
Kumar KK, O'Brien ES, Habrian CH, Latorraca NR, Wang H, Tuneew I, Montabana E, Marqusee S, Hilger D, Isacoff EY, Mathiesen JM, Kobilka BK. (2023) Negative allosteric modulation of the glucagon receptor by RAMP2. Cell. DOI: doi.org/10.1016/j.cell.2023.02.028 [link] [bioRxiv]
Silva RP, Huang Y, Nguyen AW, Hsieh CL, Olaluwoye OS, Kaoud TS, Wilen RE, Qerqez AN, Park JG, Khalil AM, Azouz LR, Le KC, Bohanon AL, DiVenere AM, Liu Y, Lee AG, Amengor DA, Shoemaker SR, Costello SM, Padlan EA, Marqusee S, Martinez-Sobrido L, Dalby KN, D'Arcy S, McLellan JS, Maynard JA. (2023) Identification of a conserved S2 epitope present on spike proteins from all highly pathogenic coronaviruses. eLife. DOI: doi.org/10.7554/eLife.83710 [link] [bioRxiv]
Glasgow A, Hobbs HT, Perry ZR, Wells ML, Marqusee S, Kortemme T. (2023) Ligand-specific changes in conformational flexibility mediate long-range allostery in the lac repressor. Nat. Commun. DOI: doi.org/10.1038/s41467-023-36798-1 [link] [bioRxiv]
Harman JL, Reardon PN, Costello SM, Warren GD, Phillips SR, Connor PJ, Marqusee S, Harms MJ. (2022) Evolution avoids a pathological stabilizing interaction in the immune protein S100A9. PNAS. DOI: doi.org/10.1073/pnas.2208029119 [link] [bioRxiv]
Hobbs HT, Shah NH, Shoemaker SR, Amacher JF, Marqusee S, Kuriyan J. (2022) Saturation mutagenesis of a predicted ancestral Syk-family kinase. Protein Science. DOI: doi.org/10.1002/pro.4411 [link] [bioRxiv]
Rosi M, Russell B, Kristensen LG, Farquhar ER, Jain R, Abel D, Sullivan M, Costello SM, Dominguez-Martin MA, Chen Y, Marqusee S, Petzold CJ, Kerfeld CA, DePonte DP, Farahmand F, Gupta S,Ralston CY. (2022) An automated liquid jet for fluorescence dosimetry and microsecond radiolytic labeling of proteins. Communications Biology. DOI: doi.org/10.1038/s42003-022-03775-1 [link]
Hidalgo F, Nocka LM, Shah NH, Gorday K, Latoracca NR, Bandaru P, Templeton S, Lee D, Karandur D, Pelton JG, Marqusee S, Wemmer DE, Kuriyan J. (2022) A saturation-mutagenesis analysis of the interplay between stability and activation in Ras. eLife. DOI: doi.org/10.7554/eLife.76595 [link] [bioRxiv]
Carroll EC, Marqusee S. (2022) Site-specific ubiquitination: Deconstructing the degradation tag. Curr Opin Struct Biol. DOI: doi.org/10.1016/j.sbi.2022.102345 [link]
Costello SM, Shoemaker SR, Hobbs HT, Nguyen AW, Hsieh CL, Maynard JA, McLellan JS, Pak JE, Marqusee S. (2022) The SARS-CoV-2 spike reversibly samples an open-trimer conformation exposing novel epitopes. NSMB. DOI: doi.org/10.1038/s41594-022-00735-5 [link] [bioRxiv]
Shaw A, Satija R, Antunez de Mayolo E, Marqusee S, Bustamante C. (2021) Rigid DNA nanotube tethers suppress high frequency noise in dual-trap optical tweezers systems. [bioRxiv]
Hobbs HT, Shah NH, Badroos JM, Gee CL, Marqusee S, Kuriyan J. (2021) Differences in the dynamics of the tandem-SH2 modules of the Syk and ZAP-70 tyrosine kinases. Protein Science. DOI: doi.org/10.1002/pro.4199 [link] [bioRxiv]
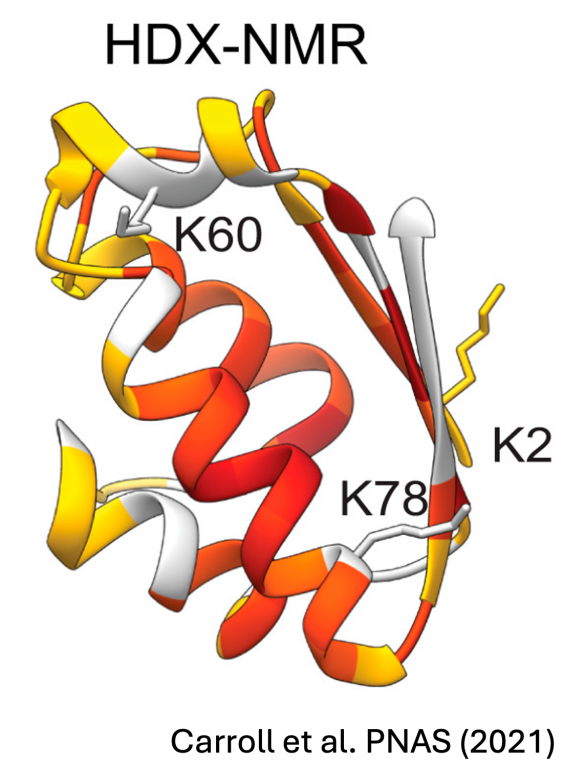
Carroll EC, Latorraca NR, Lindner JM, Maguire BC, Pelton JG, Marqusee S. (2021) Mechanistic basis for ubiquitin modulation of a protein energy landscape. PNAS. DOI: doi.org/10.1073/pnas.2025126118 [link] [bioRxiv]
Nixon CF, Lim SA, Sailer ZR, Zheludev IN, Gee CL, Kelch BA, Harms MJ, Marqusee S. (2021) Exploring the evolutionary history of kinetic stability in the α-lytic protease family. Biochemistry. DOI: doi.org/10.1021/acs.biochem.0c00720 [link] [bioRxiv]
Chen Z, Shaw A, Wilson H, Woringer M, Darzacq X, Marqusee S, Wang Q, Bustamante C. (2020) Single-molecule diffusometry reveals no catalysis-induced diffusion enhancement of alkaline phosphatase as proposed by FCS experiments. PNAS. DOI: doi.org/10.1073/pnas.2006900117 [link] [bioRxiv]
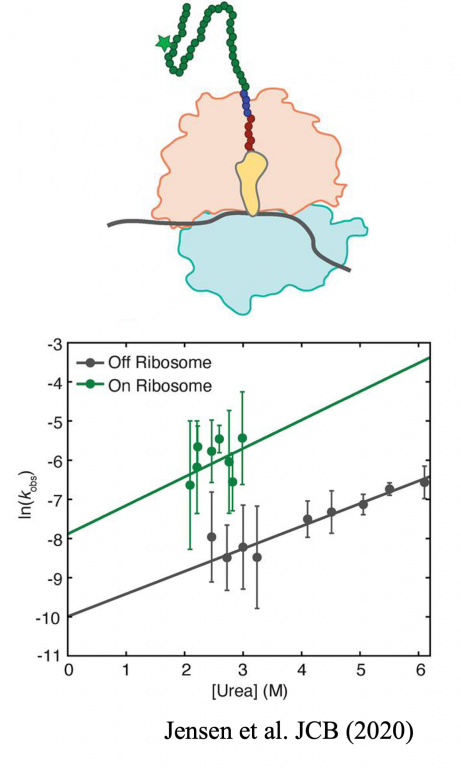
Jensen MK, Samelson AJ, Steward A, Clarke J, Marqusee S. (2020) The folding and unfolding behavior of ribonuclease H on the ribosome. JBC. DOI: 10.1074/jbc.RA120.013909 [link] [bioRxiv]
Carroll EC, Greene ER, Martin A, Marqusee S. (2020) Site-specific ubiquitination affects protein energetics and proteasomal degradation. Nat. Chem. Biol. DOI: doi.org/10.1038/s41589-020-0556-3. [link] [bioRxiv]
Oltrogge LM, Chaijarasphong T, Chen AW, Bolin ER, Marqusee S, Savage DF. (2020) Multivalent interactions between CsoS2 and Rubisco mediate α-carboxysome formation. NSMB. DOI: 10.1038/s41594-020-0387-7 [link]
Guinn EJ, Marqusee S. (2019) Using Single-Molecule Chemo-Mechanical Unfolding to Simultaneously Probe Multiple Structural Parameters in Protein Folding. Methods Protoc. DOI: 10.3390/mps2020032 [link]
Guinn EJ, Tian P, Shin M, Best RB, Marqusee S. (2018) A small single-domain protein folds through the same pathway on and off the ribosome. PNAS. DOI: 10.1073/pnas.1810517115 [link]
Rosemond SR, Hamadani KM, Cate JHD, Marqusee S. (2018) Modulating long‐range energetics via helix stabilization: a case study using T4 lysozyme. Protein Science. DOI: 10.1002/pro.3521 [link]
Lim SA, Bolin E, Marqusee S. (2018) Tracing a protein's folding pathway over evolutionary time using ancestral sequence reconstruction and hydrogen exchange. eLife. DOI: 10.7554/eLife.38369 [link]
Samelson AJ, Bolin E, Costello SM, Sharma AK, O'Brien EP, Marqusee S. (2018) Kinetic and structural comparison of a protein’s cotranslational folding and refolding pathways. Science Advances. DOI: 10.1126/sciadv.aas9098 [link]
Lim SA, Marqusee S. (2017) The burst-phase folding intermediate of ribonuclease H changes conformation over evolutionary history. Biopolymers. e23086 [link]
Guinn EJ, Marqusee S. (2017) Exploring the denatured state ensemble by single-molecule chemo-mechanical unfolding: The effect of force, temperature, and urea. J. Mol. Biol. pii: S0022-2836(17)30372-8 [link]
Hamadani KM, Howe J, Jensen MK, Wu P, Cate JHD, Marqusee S. (2017) An in-vitro tag-and-modify protein sample generation method for singlemolecule FRET. J. Biol. Chem. pii: jbc.M117.791723 [link]
Samelson AJ, Jensen MK, Soto RA, Cate JH, Marqusee S. (2016) Quantitative determination of ribosome nascent chain stability. Proc. Natl. Acad. Sci. USA 113(47):13402-13407 [link]
Lim SA, Hart KM, Harms MJ, Marqusee S. (2016) Evolutionary trend towards kinetic stability in the folding trajectory of RNases H. Proc. Natl. Acad. Sci. USA 113(46):13045-13050 [link]
Wheeler LC*, Lim SA*, Harms MJ, Marqusee S. (2016) The thermostability and specificity of ancient proteins. Curr. Opin. Structu. Biol. 38:3-43 [link] *co-first authors
Zhuravlev PI, Hinczewski M, Chakrabarti S, Marqusee S, Thirumalai D. (2016) Force-dependent switch in protein unfolding pathways and transition-state movements. Proc. Natl. Acad. Sci. USA 113(6):E715-24 [link]
Koulechova DA, Tripp KW, Horner G, Marqusee S. (2015) When the scaffold can't be ignored: The role of the hydrophobic core in ligand binding and specificity. J. Mol. Biol. doi: 10.1016/j.jmb.2015.08.014 [link]
Petzold C, Marceau AH, Miller KH, Marqusee S, Kech JL. (2015) Interaction with Single-stranded DNA-binding Protein Stimulates Escherichia coli Ribonuclease HI Enzymatic Activity. J. Biol. Chem. 290(23):14626-36 [link]
Guinn EJ, Jagannathan B, Marqusee S. (2015) Single-molecule chemo-mechanical unfolding reveals multiple transition state barriers in a small single-domain protein. Nature Communications 6:6861 [link]
Rosen LE, Marqusee S. (2015) Autonomously folding protein fragments reveal differences in the energy landscapes of homologus RNases H. PLoS One 10(3):e0119640 [link]
Bowman GR, Bolin ER, Hart KM, Maguire BC, Marqusee S. (2015) Discovery of multiple hidden allosteric sites by combining Markov State models and experiments. Proc. Natl. Acad. Sci. USA 112(9):2734-39 [link]
Riedel C, Gabizon R, Wilson CA, Hamadani K, Tsekouras K, Marqusee S, Pressé S, Bustamante C. (2015) The heat released during catalytic turnover enhances the diffusion of an enzyme. Nature.517(7533):227-30 [link] [highlight]
Hart KM, Harms MJ, Schmidt BH, Elya C, Thornton JW, Marqusee S. (2014) Thermodynamic System Drift in Protein Evolution. PloS Biology 12(11):e1001994 [link] [synopsis]
Rosen LE, Kathuria SV, Matthews CR, Bilsel O, Marqusee S. (2014) Non-Native Structure Appears in Microseconds during the Folding of E. coli RNase H. J. Mol. Biol. doi:10.1016/j.jmb.2014.10.003 [link]
Rosen LE, Connell KB, Marqusee S. (2014) Evidence for close side-chain packing in an early protein folding intermediate previously assumed to be a molten globule. Proc. Natl. Acad. Sci. USA doi:10.1073/pnas. 1410630111 [link]
Pressé S, Peterson J, Lee J, Elms P, MacCallum JL, Marqusee S, Bustamante C, Dill K. (2014) Single molecule conformational memory extraction: p5ab RNA hairpin. J. Phys. Chem. B. 118(24):6597-603. [link]
Jha SK, Marqusee S. (2014) Kinetic evidence for a two-stage mechanism of protein denaturation by guanidinium chloride. Proc. Natl. Acad. Sci. USA 111(13):4856-61. [link]
Dimster-Denk D, Tripp KW, Marini NJ, Marqusee S, Rine J. (2014) Mono and dual cofactor dependence of human cystathionine beta-synthase enzyme variants in vivo and in vitro. G3 (Bethesda) 3(10):1619-28. [link]
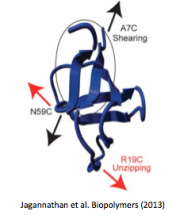
Jagannathan B, Marqusee S. (2013) Protein folding and unfolding under force. Biopolymers 99(11):860-9. [link]
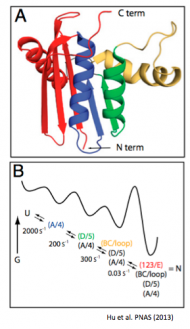
Hu W, Walters BT, Kan ZY, Mayne L, Rosen LE, Marqusee S, Englander SW. (2013) Stepwise protein folding at near amino acid resolution by hydrogen exchange and mass spectrometry. Proc. Natl. Acad. Sci. USA 110(19):7684-9. [link]
Udgaonkar J, Marqusee S. (2013) Folding and Binding. Curr Opin Struct Biol. (1):1-3. [link]
Elms P, Chodera J, Bustamante C, Marqusee S. (2012) Limitations of constant-force-feedback experiments. Biophys. J. 103(7):1490-9. [link]
Jagannathan B, Elms P, Bustamante C, Marqusee S. (2012) Direct Observation of a Force-Induced Switch in the Anisotropic Mechanical Unfolding Pathway of a Protein. Proc. Natl. Acad. Sci. USA 109(44):17820-5. [link]
Elms PJ, Chodera JD, Bustamante C, Marqusee S. (2012) The molten globule state is unusually deformable under mechanical force. Proc. Natl. Acad. Sci. USA 109(10):3796-801. [link]
Miller KH, Marqusee S. (2011) Propensity for C-terminal domain swapping correlates with increased regional flexibility in the C-terminus of RNase A. Protein Sci. 10:1735-44. [link]
Cecconi C, Shank EA, Marqusee S, Bustamante C. (2011) DNA molecular handles for single-molecule protein-folding studies by optical tweezers. Methods Mol Biol. 749:255-71. [link]
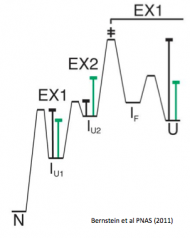
Bernstein R, Schmidt KL, Harbury PB, Marqusee S. (2011) Structural and kinetic mapping of side-chain exposure onto the protein energy landscape. Proc. Natl. Acad. Sci. USA 109(29):11681-6. [link]
Hanes MS, Ratcliff K, Marqusee S, Handel TM. (2010) Protein-protein binding affinities by pulse proteolysis: application to TEM-1/BLIP protein complexes. Protein Sci. 19(10):1996-2000. [link]
Shank EA, Cecconi C, Dill JW, Marqusee S, Bustamante C. (2010) The folding cooperativity of a protein is controlled by its chain topology. Nature 465(7298):637-40. [link]
Ratcliff K, Marqusee S. (2010) Identification of residual structure in the unfolded state of ribonuclease H1 from the moderately thermophilic Chlorobium tepidum: comparison with thermophilic and mesophilic homologues. Biochemistry 49(25):5167-75. [link]
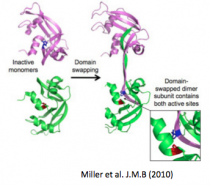
Miller KH, Karr JR, Marqusee S. (2010) A hinge region cis-proline in ribonuclease A acts as a conformational gatekeeper for C-terminal domain swapping. J. Mol. Biol. 400(3):567-78. [link]
Connell KB, Horner GA, Marqusee S. (2009) A single mutation at residue 25 populates the folding intermediate of E. coli RNase H and reveals a highly dynamic partially folded ensemble. J. Mol. Biol. 391(2):461-70. [link]
Connell KB, Miller EJ, Marqusee S. (2009) The folding trajectory of RNase H is dominated by its topology and not local stability: a protein engineering study of variants that fold via two-state and three-state mechanisms. J. Mol. Biol. 391(2):450-60. [link]
Ratcliff K, Corn J, Marqusee S. (2009) Structure, stability, and folding of ribonuclease H1 from the moderately thermophilic Chlorobium tepidum: comparison with thermophilic and mesophilic homologues. Biochemistry 48(25):5890-8. [link]
Cecconi C, Shank, E, Dahlquist FW, Marqusee S, Bustamante C. (2008) Protein-DNA chimeras for single molecule mechanical folding studies with the optical tweezers. European Biophysics Journal 37(6):729-38. [link]
Celletti J, Bernstein R, Marqusee S. (2007) Exploring Subdomain Cooperativity in T4 Lysozyme II: uncovering the C-terminal subdomain as a hidden intermediate in the kinetic folding pathway. Protein Sci. 16(5):852-62. [link]
Cellitti J, Llinas M, Echols N, Shank EA, Gillespie B, Crowder SM, Dahlquist FW, Alber T, Marqusee S. (2007) Exploring Subdomain Cooperativity in T4 Lysozyme I: structure and energetic studies of a circular permutant and protein fragment. Protein Sci. 16(5):842-51. [link]
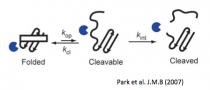
Park C., Zhou S., Gilmore J., Marqusee S. (2007) Energetics-Based Protein Profiling on a Proteomic Scale: Identification of Proteins Resistant to Proteolysis. J. Mol. Biol. 368(5):1426-37. [link]
Young TA, Skordalakes E, and Marqusee, S. (2007) Comparison of proteolytic susceptibility in phophoglycerate kinases from yeast and E. coli: modulation of conformational ensembles without altering structure or stability. J. Mol. Biol. 368(5):1438-47. [link]
Cecconi C, Shank, E, Marqusee S, Bustamante C. (2007) Studying protein folding with laser tweezers. Proceedings of the International School Enrico Fermi. [link]
Freedman TF, Sondermann H, Friedland G, Kortemme T, Bar-Sagi, Marqusee S, Kuriyan J. (2006) A Ras-induced conformational switch in the Ras activator Son of Sevenless. Proc. Natl. Acad. Sci. USA 103: 16692-16697. [link]
Park C, Marqusee S. (2006) Quantitative Determination of Protein Stability and Ligand Binding by Pulse Proteolysis. Current Protocols in Protein Science Chapter 20:Unit 20.11 [link]
Wildes, DE, Anderson M, Sabogal A, Marqusee S. (2006) Native state energetics of the Src SH2 domain: Evidence for a partially structured state in the denatured ensemble. Protein Science (1769-79). [link]
de los Rios MA, Muralidhara BK, Wildes D, Sosnick TR, Marqusee S, Wittung-Stafshede P, Plaxco KW, Ruczinski I. (2006) On the precision of experimentally determined protein folding rates and phi-values. Protein Sci. 15(3):553-63. [link]
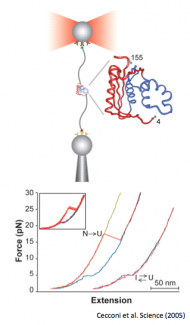
Cecconi, C, Shank E, Bustamante C, Marqusee S. (2005) Direct Observation of the three-state folding of a single protein molecule. Science 309:2057-60. [link]
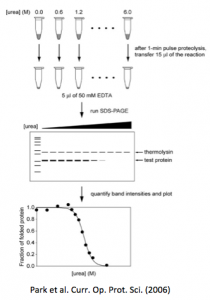
Park C, Marqusee S. (2005) Pulse Proteolysis: A Simple Method for Quantitative Determination of Protein Stability and Ligand Binding. Nature Methods 2(3):207-12. [link]
Maxwell KL, Wildes D, Zarrine-Afsar A, de los Rios MA, Brown AG, Friel CT, Hedberg L, Horng J-C, Bona D, Miller EJ, Valle-Blisle A, Main ERG, Bemporad F, Qiu L, Teilum K, Vu N-D, Edwards AM, Ruczinski I, Poulsen FM, Kragelund BB, Michnick SW, Chiti F, Bai Y, Hagen SJ, Serrano L, Oliveberg M, Raleigh DP, Wittung-Stafshede P, Radford SE, Jackson SE, Sosnick TR, Marqusee S, Davidson AR, Plaxco KW. (2005) Protein folding: defining a standard set of experimental conditions and a preliminary kinetic data set of two-state proteins. Protein Sci. 14(3):602-16. [link]
Wildes D, Marqusee S. (2005) Hydrogen exchange and ligand binding: Ligand-dependent and ligand-independent protection in the Src SH3 domain. Protein Sci. 14(1):81-8. [link]
Park C, Marqusee S. (2004) Probing the High Energy States in Proteins by Proteolysis. J. Mol. Biol. 345:1467–76. [link]
Park C, Marqusee S. (2004) Analysis of the stability of multimeric proteins by effective DG and effective m-values. Protein Sci. 13:2553-8. [link]
Wildes D, Marqusee S. (2004) Hydrogen Exchange Strategies Applied to Protein Ensemble. Methods in Enzymology 380:328-49. [link]
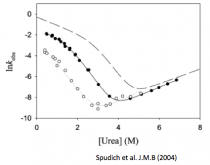
Spudich GM, Miller EJ, Marqusee S. (2004) Destabilization of the Escherichia coli RNase H kinetic intermediate: switching between a two-state and three-state folding mechanism. J. Mol. Biol. 335(2):609-18. [link]
Robic S, Guzman-Casado M, Sanchez-Ruiz JM, Marqusee S. (2003) Role of residual structure in the unfolded state of a thermophilic protein. Proc. Natl. Acad. Sci. USA 100:11345-9. [link]
Guzman-Casado M, Parody-Morreale A, Robic S, Marqusee S, Sanchez-Ruiz JM. (2003) Energetic evidence for formation of a pH-dependent hydrophobic cluster in the denatured state of Thermus thermophilus ribonuclease H. J. Mol. Biol. 329(4):731-43. [link]
Kim R, Lai L, Lee HH, Cheong GW, Kim KK, Wu Z, Yokota H, Marqusee S, Kim SH. (2003) On the mechanism of chaperone activity of the small heat-shock protein of Methanococcus jannaschii. Proc. Natl. Acad. Sci. USA 100:8151-5. [link]
Kern G, Pelton J, Marqusee S, Kern D. (2002) Structural Properties of the histidine-containing loop in HIV-1 RNase H. Biophys. Chem. 96:285-91. [link]
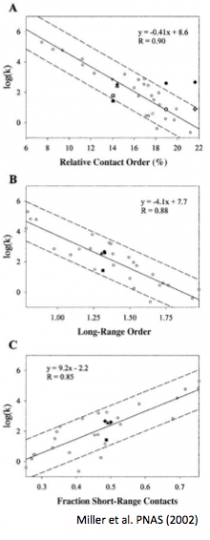
Miller EJ, Fischer KF, Marqusee S. (2002) Experimental evaluation of topological parameters determining protein-folding rates. Proc. Natl. Acad. Sci. USA 99(16):10359-63. [link]
Nicholson E, Mo H, Cohen FE, Prusiner SB, Marqusee S. (2001) Differences between the prion protein and its homolog Dpl: a partially structured state with implications for scrapie formation. J. Mol. Biol. 316(3):807-15. [link]
Spudich G, Lorenz S, Marqusee S. (2002) Propagation of a Single Destabilizing Mutation throughout the E. coli Ribonuclease HI Native State. Protein Sci.11:522-8. [link]
Hollien J, Marqusee S. (2002) Comparison of the folding processes of T. thermophilus and E. coli Ribonucleases H. J. Mol. Biol. 316:327-40. [link]
Robic S, Berger JM, Marqusee S. (2002) Contributions of the folding cores to thermostability of two Ribonucleases H. Protein Sci. 11:381-9. [link]
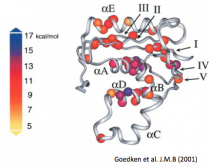
Goedken ER, Marqusee S. (2001) Native-state energetics of a thermostabilized variant of ribonuclease H. J. Mol. Biol. 314:863-71. [link]
Goedken ER, Marqusee S. (2001) Co-crystal of E. coli RNase HI with MN2+ reveals two divalent metals bound in the active site. J. Biol. Chem. 276: 7266-71. (cover article) [link]
Parker MJ, Marqusee S. (2001) A kinetic folding intermediate probed by native state exchange. J. Mol. Biol. 305:593-602. [link]
Goedken, ER, Keck JL, Berger JM, Marqusee S. (2000) Divalent metal cofactor binding in the kinetic folding trajectory of E. coli ribonuclease HI. Protein Sci. 9:1914-21. [link]
Spudich G, Marqusee S. (2000) A change in the apparent m value reveals a populated intermediate under equilibrium conditions in E. coli ribonuclease HI. Biochemistry 39:11677-83. [link]
Fischer KF, Marqusee S. (2000) A rapid test for identification of autonomous folding units in proteins. J. Mol. Biol. 302:701-712. [link]
Toth EA, Worby C, Dixon JE, Goedken ER, Marqusee S, Yates TO. (2000) The crystal structure of adenylosuccinate lyase from Pyrobaculum aerophilum reveals an intracellular protein with three disulfide bonds. J. Mol. Biol. 301:433-450. [link]
Parker MJ, Marqusee S. (2000) A Statistical Appraisal of Native State Hydrogen Exchange Data: Evidence for a Burst Phase Continuum? J. Mol. Biol. 300:1361-1375. [link]
Parker MJ, Marqusee S. (1999) The Cooperativity of Burst Phase Reactions Explored. J. Mol. Biol. 293:1195-210. [link]
Hollien J, Marqusee S. (1999) Structural distribution of stability in a thermophilic enzyme. Proc. Natl. Acad. Sci. USA 96:13674-8. [link]
Chamberlain AK, Marqusee S. (1999) Comparison of equilibrium and kinetic approaches for determining protein folding mechanisms. Advances in Protein Chemistry 53:283-328, Edited by R.C. Matthews. [link]
Llinas M, Gilespie B, Dahlquist FW, Marqusee, S. (1999) The energetics of T4 lysozyme reveal a hierarchy of conformations. Nature Structural Biology 6:1072-8. [link]
Chamberlain AK, Fischer KF, Reardon D, Handel TM, Marqusee S. (1999) Folding of an isolated ribonuclease H core fragment. Protein Sci. 8:2251-7. [link]
Goedken ER, Marqusee S. (1999) Metal binding and activation of the ribonuclease H domain from moloney murine leukemia virus. Protein Engineering 12:975-80. [link]
Raschke TM, Kho J, Marqusee S. (1999) Confirmation of the hierarchical folding of RNase H: a protein engineering study. Nature Structural Biology 6:825-831. [link]
Liu H, Farr-Jones S, Ulyanov N, Llinas M, Marqusee S, Cohen FE, Prusiner SB, James TL. (1999) Solution structure of Syrian hamster prion protein rPrP(90-231). Biochemistry 38:5362-77. [link]
Hollien J, Marqusee S. (1999) A thermodynamic comparison of mesophilic and thermophilic ribonucleases H. Biochemistry 38:3831-6. [link]
Goedken ER, Marqusee S. (1998) Ribonuclease H. Encyclopedia of Molecular Biology, Edited by T.E. Creighton.
Keck JL, Goedken ER, Marqusee S. (1998) Activation/attenuation model for RNase H. A one-metal mechanism with second-metal inhibition. J. Biol. Chem. 273:34128-33. [link]
Goedken ER, Marqusee S. (1998) Folding the ribonuclease H domain of Moloney murine leukemia virus reverse transcriptase requires metal binding or a short N-terminal extension. Proteins 33:135-43. [link]
Raschke TM, Marqusee S. (1998) Hydrogen exchange studies of protein folding. Current Opinions in Biotechnology 9:80-6. [link]
Kern G, Handel TM, Marqusee S. (1998) Characterization of a folding intermediate from HIV-1 RNase H. Protein Sci. 7:2164-2174. [link]
Chamberlain AK, Marqusee S. (1998) Molten globule energetics monitored by hydrogen exchange in urea. Biochemistry 37:1736-1742. [link]
Llinas M, Marqusee S. (1998) Subdomain interactions as a determinant in the folding and stability of T4 Lysozyme. Protein Sci. 7:96-104. [link]
Chamberlain AK, Marqusee S. (1997) Touring the landscapes: partially folded proteins examined by hydrogen exchange. Structure 5:859-863. [link]
Raschke TM, Marqusee S. (1997) The kinetic folding intermediate of ribonuclease H resembles the acid molten globule and partially unfolded molecules detected under native conditions. Nature Structural Biology 4:298-304. [link]
Goedken ER, Raschke TM, Marqusee S. (1997) Importance of the C-terminal helix to the stability and enzymatic activity of E. coli ribonuclease H. Biochemistry 36:7256-63. [link]
Chamberlain AK, Handel TM, Marqusee S. (1997) Detection of protein unfolding and fluctuations by native state hydrogen exchange. Techniques in Protein Chemistry VIII. Edited by Daniel Marshak. [link]
Keck JL, Marqusee S. (1997) Metal activation and regulation of E. coli RNase H. Techniques in Protein Chemistry VIII, Edited by Daniel Marshak .
Dabora JM, Pelton JG, Marqusee S. (1996) Structure of the acid state of E. coli Ribonuclease HI. Biochemistry 35:11951-8. [link]
Chamberlain AK, Handel TM, Marqusee S. (1996) Detection of rare partially folded molecules in equilibrium with the native conformations of RNase H. Nature Structural Biology 3:782-7. [link]
Keck JL, Marqusee S. (1996) The putative substrate recognition loop of E. coli ribonuclease H is not essential for activity. J. Biol. Chem. 271:19883-7. [link]
Keck JL, Marqusee S. (1995) Substitution of a highly basic helix/loop sequence into the HIV RNase H domain restores its Mn++-dependent RNase H activity. Proc. Natl. Acad. Sci. USA 7:2740-4. [link]
Dabora JM, Marqusee S. (1994) Equilibrium unfolding of E. coli ribonuclease H: characterization of a partly folded state. Protein Sci. 3:1401-8. [link]